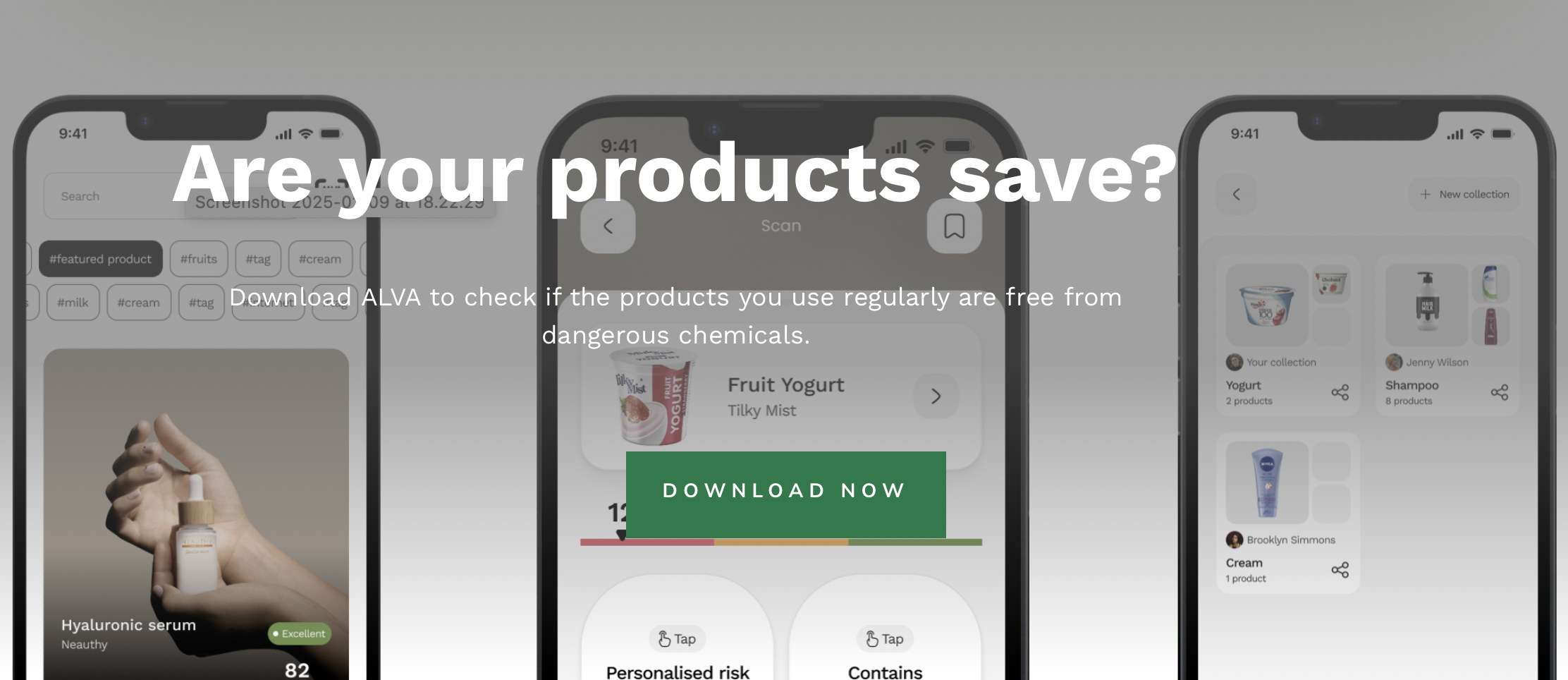
Alva is a product risk scanner that helps people understand what is actually in the things they buy. The iOS app handles scanning and presentation, but the hard problem lives in the backend: how do you take a raw ingredient list from a barcode scan and turn it into a meaningful, personalized risk assessment fast enough for someone standing in a store?
This post covers the backend architecture, the chemical detection pipeline, how the multi-factor risk scoring works, and the trade-offs we made along the way.
The stack
The backend runs on Node.js with Express, PostgreSQL with the pgvector extension for vector similarity search, and OpenAI for both embeddings and analysis. It is deployed on Render with Docker. There is no separate ML service or microservice split — the whole system runs in one process with async job processing handled internally.
That simplicity was intentional. At the current scale of over 2,300 users, the operational cost of a distributed architecture would not be justified by the performance gains. When a scan comes in, the work is I/O-bound — external API calls and database queries — not CPU-bound. A single process with async jobs handles that well.
What happens when a product is scanned
A barcode scan triggers a pipeline that looks roughly like this:
- The client sends the UPC code
- The backend checks whether the product already exists in the database — if it does, it returns the cached result immediately
- If the product is new, the backend creates a processing job and returns a job ID so the client can poll for status
- The UPC is looked up via an external product API to get product metadata and the raw ingredient list
- If the ingredients are not in English, they are translated using OpenAI
- The ingredient string is split and each ingredient is run through the chemical detection pipeline
- Risk scoring is calculated
- The product is stored with its full analysis and a push notification is sent to the user
Steps 3 through 8 happen asynchronously. The client polls the job status endpoint until the result is ready. That matters because a full analysis involves multiple external calls and can take several seconds — making the user stare at a loading screen on a synchronous request would break the experience.
The chemical detection problem
The core technical challenge is: given an ingredient name like “cetearyl alcohol” or “sodium lauryl sulfate,” determine whether it matches a known harmful chemical.
This sounds like a string-matching problem, but it is not. Ingredient lists are inconsistent. The same chemical appears under different names, abbreviations, IUPAC notations, or trade names depending on the product, the market, and the labeling conventions. Exact string matching misses too much.
How we solved it with vector search
Instead of maintaining a lookup table of name variants, we embedded each chemical in the database as a 1536-dimensional vector using OpenAI’s text-embedding-3-small model. The chemical table holds EPA DTXSID records with preferred names, IUPAC names, CASRN identifiers, InChI keys, molecular formulas, and ToxCast toxicity data.
When an ingredient comes in, we generate its embedding and run a cosine similarity search against the chemical table using pgvector. Any match above a 0.6 similarity threshold is flagged as a potential chemical match.
This approach has three properties we needed:
- Resilience to naming variation. “Perfluorooctanoic acid” and “PFOA” both land close to the same vector neighborhood. So do misspellings and partial names.
- Semantic matching, not keyword matching. The embedding captures meaning, so structurally related chemicals cluster together even if the names share no common substrings.
- Low maintenance. Adding new chemicals to the database requires generating an embedding, not manually curating alias tables.
The trade-off is that vector search is fuzzier than exact matching. A 0.6 threshold sometimes catches ingredients that are structurally similar but not actually harmful. That is why vector matching is just the first layer — the AI scoring layer provides the nuance.
PFAS detection specifically
PFAS — per- and polyfluoroalkyl substances — are one of the main concerns Alva users care about. These are the “forever chemicals” that accumulate in the body and appear in everything from food packaging to cosmetics.
The detection approach is the same vector pipeline, but the chemical database includes the specific PFAS family of compounds. Because vector search captures structural relationships, it catches PFAS-adjacent compounds that a keyword search for “PFAS” or “PFOA” would miss entirely.
Once a product’s ingredients have been matched against the chemical database, the matched chemical names are stored in a dedicated array on the product record. The count of matches provides a preliminary risk classification:
- Zero matches: low risk
- One match: high risk
- Two or more: very high risk
That simple classification is useful for quick scanning, but the product needed more than a traffic light. Users wanted to understand why a product was flagged and what it meant for them personally.
Multi-factor risk scoring
The deeper risk assessment runs four parallel AI analyses and combines them with configurable weights:
Toxicity analysis (40% weight)
This is the heaviest factor. The system sends the product’s ingredient list and matched chemical data to OpenAI and asks for a toxicity score from 1 to 10, along with a toxicity level classification and explanation. The prompt includes the specific chemicals found and asks the model to evaluate their known health effects.
Ingredient concentration analysis (30% weight)
A chemical being present matters less if it is the last ingredient on a list of fifty. This analysis estimates how concentrated the problematic ingredients are likely to be, based on their position in the ingredient list, the product type, and typical formulation patterns. Ingredient lists are ordered by concentration, which gives the model meaningful signal.
Exposure analysis (20% weight)
A face cream you apply daily is a different risk profile than a cleaning product you use once a month. This analysis evaluates how frequently and for how long a user is likely exposed to the product based on its category and typical usage patterns.
Health risk analysis (10% weight)
This is where personalization happens. During onboarding, users provide their health conditions, concerns, age, and the specific risks they care about — reproductive health, cancer risk, immune concerns, endocrine disruption. The health risk analysis takes the product’s chemical profile and scores it against the user’s specific health data.
Combined score
The four scores are combined with their weights into a total risk score from 0 to 10:
totalRiskScore = (0.4 × toxicity) + (0.3 × concentration) + (0.2 × exposure) + (0.1 × health_risk)The full result — including each sub-analysis with its score and explanation — is stored as a JSONB object on the product record. That means the iOS app can show the total score as a headline number and let users drill into each factor if they want the detail.
Why JSONB for risk data
We store risk analysis results, chemical search results, and user profile data as JSONB rather than normalized relational tables.
The reason is practical. The shape of the risk analysis evolved several times during development as we tuned the prompts, adjusted the weight factors, and added the personalization layer. With JSONB, those changes did not require database migrations. The trade-off is that you lose relational integrity and query efficiency on nested fields, but for data that is written once and read as a whole document, that trade-off is worth it.
Per-user risk caching
The personalized risk score depends on the user’s health profile, which means the same product can have different risk scores for different users. Computing that on every request would be wasteful, so we cache the per-user risk assessment in a dedicated table.
When a user requests risk data for a product, the backend checks the cache first. If there is no cached result and the user has health concerns on file, the full multi-factor analysis runs and the result is stored. If the user later updates their health profile, the cached results can be invalidated.
Push notifications for async results
Because product analysis is async, the user needs to know when their result is ready. The backend sends a push notification via Firebase Cloud Messaging as soon as the analysis completes. The iOS app handles this with standard APNs delivery, and the notification deep-links back to the product result screen.
This solved a real UX problem. Early versions relied entirely on polling, which meant users had to keep the app open and wait. Push notifications let them scan, put their phone away, and come back when the result is ready.
The biggest challenges
Embedding quality versus speed
We chose text-embedding-3-small for its speed and cost. The 1536-dimensional vectors are good enough for chemical name matching, but they occasionally surface false positives — ingredients that are structurally similar in embedding space but not actually harmful. The AI scoring layer compensates for this, but it means the system works best as a pipeline rather than relying on any single stage.
Prompt stability
The risk scoring depends on OpenAI returning structured JSON with consistent scoring behavior. In practice, prompt engineering for consistent numerical scoring is harder than it looks. Small changes in wording can shift the distribution of scores. We settled on explicit scoring criteria in each prompt and JSON mode to enforce the response structure.
Ingredient list quality
Real-world ingredient lists are messy. They come in different languages, use inconsistent delimiters, include parenthetical sub-ingredients, and sometimes arrive as images rather than text. The translation and parsing step before chemical matching has to be forgiving enough to handle that variation without silently dropping ingredients.
What we would do differently
If we were starting today, we would probably separate the embedding generation into a background job that pre-computes and caches ingredient embeddings rather than generating them on the fly during a scan. That would reduce latency for products with many ingredients and lower the per-scan cost of OpenAI API calls.
We would also consider a hybrid matching approach: exact string matching against known ingredient names first, with vector search as a fallback for unrecognized ingredients. That would give us the speed of lookups for common ingredients and the resilience of semantic search for edge cases.
Why this architecture works for Alva
The system works because it treats chemical detection as a layered problem rather than a binary lookup. Vector search provides broad, resilient matching. AI scoring adds contextual nuance. Health personalization makes the result actionable for an individual user. And async processing keeps the scanning experience responsive even when the analysis takes time.
At 2,300 users and growing, the architecture handles the current load comfortably. The next scaling challenges will be embedding cache efficiency, prompt cost optimization, and expanding the chemical database as new research surfaces. But the layered approach gives us room to improve each stage independently without rearchitecting the whole pipeline.